One advantage of using XML is that one can rely on good support in libraries for functionality. When parsing XML, one does not have to take care of the syntax, and focus on the data and its semantics. This comes at the expense of verbosity, though, but having the ability to express semantics explicitly is a huge benefit for flexibility.
So, when
Peter and Henry put their first documents online about the Chemical Markup Language (CML), I was thrilled, even though is actually was still SGML when I encountered it. The work predates the
XML recommendation. As I
recently blogged, in '99 I wrote patches for Jmol and JChemPaint to support CML, which were published as preprint in the
Chemical Preprint Server in a paper in 2000 in the
Internet Journal of Chemistry. Neither of the two has survived.
Anyway, the
Chemistry Development Kit makes heavy use of CML, and
Bioclipse supports it too. Now, Bioclipse is based on the
Eclipse Rich Client Platform architecture, for which there exist quite a few XML tools in the
Web Tools Platform (WTP). Among these, a validation, content assisting XML editor. This means, I get red markings when I make my XML document not-well-formed or invalid. Just a quick recap: well-formedness means that the XML document has a proper syntax: one root node, properly closed tags, quotes around attribute values, etc. Validness, however, means that the document is well-formed, but also hierarchically organized according to some specification.
Enter CML. CML is such a specification, first with DTDs, but after the introduction of XML Namespaces with XML Schema (see
There can be only one (namespace)). The WTP can use this XML Schema for validation, and this is of great help learning the CML language. Pressing Ctrl-space in Bioclipse will now show you what allowed content can be added at the current character position.
Yes, Bioclipse can do this now (in SVN, at least). This has been on my wishlist for at least two years now, but never really found the right information. Now, three days ago
David wrote about
End of Year Cramps in which he describes some of his work on the WTP for autocomplete for XPath queries. He
see[s] a brighter future for XML at eclipse over the next year. I hope that those in the eclipse and XML community will help to continue to improve the basic support, so that first class commercial quality applications that leverage this support can continue to be built.That was enough statement for me to
ask in the comments on how to make the WTP XML editor aware of the CML XML Schema. It already picked up XML Schema's with
xsi:schemaLocation, but I needed something to worked without such statements in the XML document itself. David explained that me that I could use the
org.eclipse.wst.xml.catalog extension. This was really easy, and
commited to Bioclipse SVN as:
<extension
point="org.eclipse.wst.xml.core.catalogContributions">
<catalogContribution>
<uri name="http://www.xml-cml.org/schema"
uri="schema24/schema.xsd"/>
</catalogContribution>
</extension>
However, that does not make the WTP XML editor available in the Bioclipse application yet. Not ever in the "Open With"... So, I set up a
CML Feature. After a follow up question, it turned out that the CML content type of Bioclipse was already a sub type of the XML type (see ):
<extension
point="org.eclipse.core.runtime.contentTypes">
<content-type
base-type="org.eclipse.core.runtime.xml"
id="net.bioclipse.contenttypes.cml"
name="Chemical Markup Language (CML)"
file-extensions="cml,xml"
priority="normal">
</content-type>
</extension>
So, the only remaining problem was to actually get the WTP XML editor as part of the Bioclipse application. The new CML Feature takes care of that (I hope the export and building the update site work too, but that's yet untested), by important the relevant plugins and features. Last night, however, I ended up with one stacktrace which gave me little clue on which plugin I was still missing.
Therefore, I headed to #eclipse and actually met David of the blog that started this again. He asked
nitind to think about it too, and they helped me pin down the issue. This relevant bit of the stacktrace turned out to be:
Caused by: java.lang.IllegalStateException
at org.eclipse.core.runtime.Platform.getPluginRegistry(Platform.java:774)
at org.eclipse.wst.common.componentcore.internal.impl.WTPResourceFactoryRegistry$ResourceFactoryRegistryReader.(WTPResourceFactoryRegistry.java:275)
at org.eclipse.wst.common.componentcore.internal.impl.WTPResourceFactoryRegistry.(WTPResourceFactoryRegistry.java:61)
at org.eclipse.wst.common.componentcore.internal.impl.WTPResourceFactoryRegistry.(WTPResourceFactoryRegistry.java:55)
... 37 more
This refered to this bit of code of Eclipse' Platform.java:
Bundle compatibility = InternalPlatform.getDefault()
.getBundle(CompatibilityHelper.PI_RUNTIME_COMPATIBILITY);
if (compatibility == null)
throw new IllegalStateException();
So, the plugin I turned to to have missing was
org.eclipse.core.runtime.compatibility. Apparently, some parts of the WTP that the XMLEditor is using, still uses Eclipse2.x technology.

This screenshot shows the WTP XMLEditor in action in Bioclipse on a CML file. It shows the document contents with the 'Design' tab, which also shows allowed content, as derived from the XML Schema for CML. Also, note that the Outline and Properties view automatically come for free, which allows more detail and navigation of the content.

This screenshot shows the 'Source' tab for the same file, where I deliberately changed the value of the @id attribute of the first atom. The value does not validate against the regular expression defined in the CML schema for @id attribute values. It also shows the content assisting in action. At any location in the CML file, I can hit Ctrl-Space, and the editor will show me which content I can add at that location.
This makes Bioclipse a perfect tool to craft CML documents and learn the language.
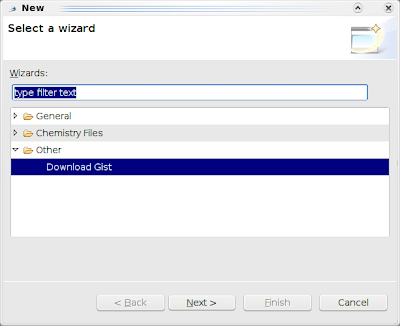 and click Next:
and click Next: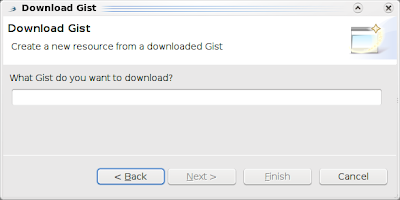 Then, just type the number of the Gist you want to open in Bioclipse, for example 18315 (see Bioclipse2 Scripting #1: from SMILES to a UFF optimized structure in Jmol), and click another Next to select a file name and location:
Then, just type the number of the Gist you want to open in Bioclipse, for example 18315 (see Bioclipse2 Scripting #1: from SMILES to a UFF optimized structure in Jmol), and click another Next to select a file name and location: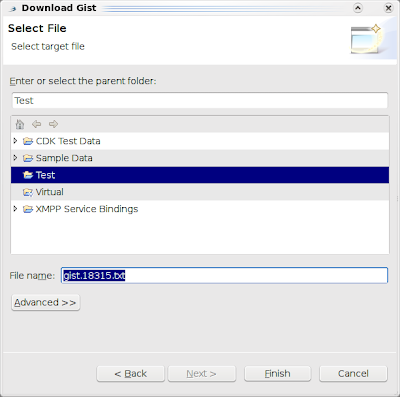 The current code does require you to know the Gist number, so you'll need a web browser to look it up, but we do have search facilities in mind. Also, while the code attempts so, the resulting Gist is not automatically openend in an editor (a bug). Another idea is to just install the egit plugin in Bioclipse :)
The current code does require you to know the Gist number, so you'll need a web browser to look it up, but we do have search facilities in mind. Also, while the code attempts so, the resulting Gist is not automatically openend in an editor (a bug). Another idea is to just install the egit plugin in Bioclipse :)


